How robotic bronchoscopy helped a Georgia man avoid unnecessary lung surgery

When Dan Hannon, 72, received guidance to have part of a lung taken out whether nodules were cancerous or not, he wanted a second opinion to see if he would indeed live the rest of his life without a full set of lungs.
After incidentally learning of a suspicious spot in his lungs while getting imaging to diagnose a kidney stone in 2025, he was advised to get his lungs checked. Several months later, he had a diagnostic PET scan under the care of a Georgia pulmonologist near his home in Buford, Georgia.
Some spots “lit up,” indicating metabolic activity, which often means cancer but may be due to infection or inflammation. He went on to get a bronchoscopy, a minimally invasive procedure using a camera on a thin tube to examine his lungs and obtain biopsy tissue.
He and his wife, Janice, learned the areas that were successfully biopsied were benign, but there were two additional areas the physician couldn’t reach.
Thus came the recommendation for an open surgery to remove the undefined tissue for biopsy. It was also advised that Hannon have a sizeable portion of his affected lung removed while still under anesthesia, regardless of the biopsy results.
That didn’t sit well with the couple, so they drove four hours to Vanderbilt Health in Nashville. Their son had been successfully treated for cancer there in the past.
“The prior lack of knowledge of whether I had cancer or not, and being told I should have surgery and get part of my lung cut out — whether it was cancer or not — was very bothersome,” Hannon said. “I decided to get a second opinion. I wanted all the information I could get to make a good decision, and that’s exactly what’s happened.”
At Vanderbilt Health, a new approach to a difficult diagnosis
The Hannons met with Fabien Maldonado, MD, MSc, professor of Medicine and Thoracic Surgery and director of Interventional Pulmonology at the Vanderbilt Lung Institute. Maldonado told them he was pretty confident he could reach those areas for biopsy using robotic bronchoscopy, the standard of care at Vanderbilt University Hospital.
“Robotic bronchoscopy, combined with cone-beam CT, has transformed the way we do things,” said Maldonado, who holds the Pierre Massion Directorship in Lung Cancer Research. “Ninety-five percent of people with lung nodules don’t have cancer, but for the 5% that do, they need to receive a diagnosis and get treatment as soon as possible.
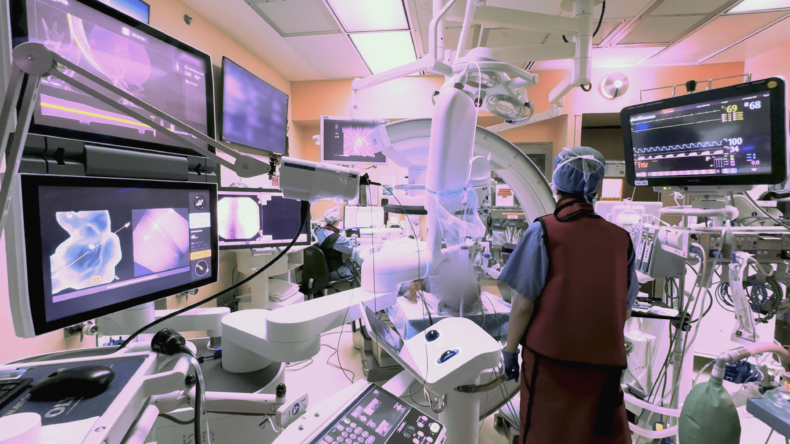
Inside the robotic bronchoscopy advancing lung care
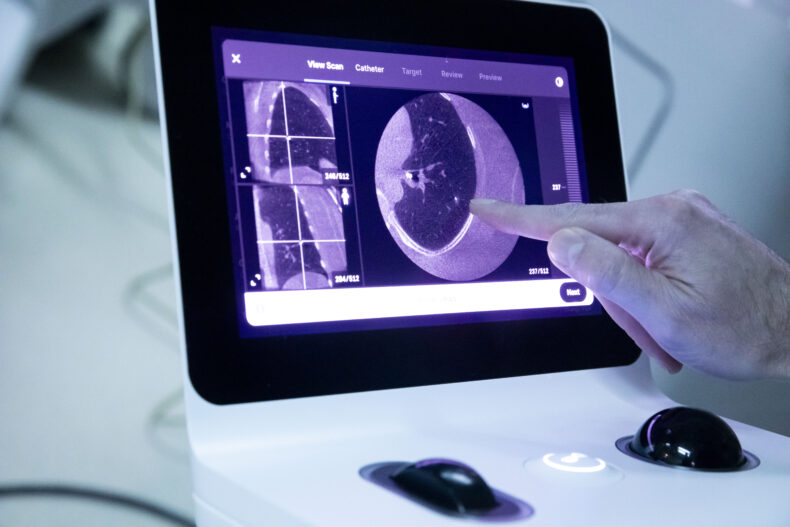
During a robotic bronchoscopy, physicians use a controller to precisely guide a bronchoscopy tube, which is typically smaller and more flexible than traditional tubes. The additional agility allows access into the lung’s harder-to-reach peripheral structures so biopsy tissue can be obtained. A 3D, high-resolution image of the lung obtained through cone-beam computed tomography (CBCT) is displayed on a monitor to guide the interventional pulmonologist’s progress.
At Vanderbilt Health, the four board-certified interventional pulmonologists who do the robotic bronchoscopies have all completed a fellowship and an additional year of interventional pulmonology training before receiving board certification from the American Association for Bronchology and Interventional Pulmonology.
Oncologist Mohamed Shanshal, MBChB, assistant professor of Medicine, said robotic bronchoscopy significantly improves getting patients the appropriate treatment as soon as possible, reducing their anxiety and improving care.
To expand access to the diagnostic procedure, the Vanderbilt Health Interventional Pulmonology program recently bought two additional robot and CBCT systems, making it one of the largest interventional pulmonology programs in the South. Four state-of-the-art bronchoscopy suites are slated to open later this year.
“This reduces diagnostic uncertainty and helps us move more quickly from suspicion to confirmed diagnosis,” Shanshal said. “Earlier and more accurate tissue diagnosis allows us to initiate treatment sooner, including surgery, targeted therapy, immunotherapy or clinical trials. In lung cancer, timing and adequate tissue for molecular testing are critical to optimizing outcomes.”
Maldonado explained that the Interventional Pulmonology program is a national leader in testing new technologies for safety and results.
“Many new technologies related to lung nodule biopsies have come on the market without any data to prove their benefit,” Maldonado said. “We are doing the randomized controlled trials here to prove whether something new is better than what we’re already doing. We study these technologies carefully and quickly, and within six months to a year, we know if the new is more beneficial and needs to be adopted.”
From uncertainty to action: What Hannon’s biopsy uncovered
For the Hannons, having the assurance that the robotic bronchoscopy would reach the spots in his lung and offer clearer details about their makeup was important.
“With the help of the robot, Dr. Maldonado was able to get what he needed, and it came back as adenocarcinoma,” Hannon said. “He was awesome when he explained his findings and everything we needed to do.”
Hannon has mucinous adenocarcinoma, a rare, non-small cell lung cancer that accounts for 2-10% of all lung adenocarcinomas and is most often found in the outer regions of the lung.
During a return visit to Nashville in late February, Hannon had a pulmonary function test, and met with Konrad Hoetzenecker, MD, PhD, professor of Thoracic Surgery, who will soon surgically remove the cancerous spots, and with Shanshal, his new oncologist.
“We’ve had a fantastic experience at Vanderbilt, and we’ve got a fantastic team of medical professionals we’re working with,” Hannon said. “We’re prepared for surgery and any future treatment because of the consultations we’ve had with Dr. Maldonado, Dr. Hoetzenecker and Dr. Shanshal. This has given me confidence in what has to happen.”
And as the Hannons move with hope toward healing, they’re looking forward to future cruises and travels to visit family.
The post How robotic bronchoscopy helped a Georgia man avoid unnecessary lung surgery appeared first on Vanderbilt Health News.






